Personalized Cancer Therapy Prioritization Based on Driver Alteration Co-occurrence Patterns
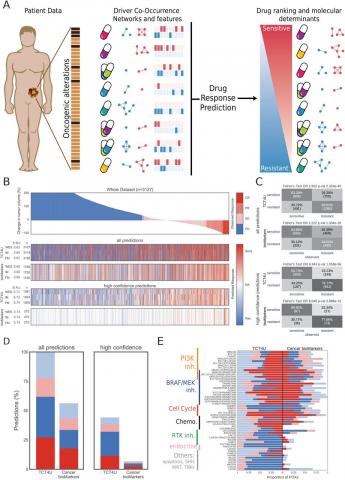
Mateo L, Duran-Frigola M, Gris-Oliver A, Palafox M, Scaltriti M, Razavi P, Chandarlapaty S, Arribas J, Bellet M, Serra V, Aloy P, Molecular profiling of personal cancer genomes, and the identification of actionable vulnerabilities and drug-response biomarkers, are the basis of precision oncology. Tumors often present several driver alterations that might be connected by cross-talk and feedback mechanisms, making it difficult to mark single oncogenic variations as reliable predictors of therapeutic outcome. In the current work, we uncover and exploit driver alteration co-occurrence patterns from a recently published in vivo screening in patient-derived xenografts (PDXs), including 187 tumors and 53 drugs. For each treatment, we compare the mutational profiles of sensitive and resistant PDXs to statistically define Driver Co-Occurrence (DCO) networks, which capture both genomic structure and putative oncogenic synergy. We then use the DCO networks to train classifiers that can prioritize, among the available options, the best possible treatment for each tumor based on its oncogenomic profile. In a cross-validation setting, our drug-response models are able to correctly predict 66% of sensitive and 77% of resistant drug-tumor pairs, based on tumor growth variation. Perhaps more interesting, our models are applicable to several tumor types and drug classes for which no biomarker has yet been described. Additionally, we experimentally validated the performance of our models on 15 new tumor samples engrafted in mice, achieving an overall accuracy of 75%. Finally, we adapted our strategy to derive drug-response models from continuous clinical outcome measures, such as progression free survival, which better represent the data acquired during routine clinical practice and in clinical trials. We believe that the computational framework presented here could be incorporated into the design of adaptive clinical trials, revealing unexpected connections between oncogenic alterations and increasing the clinical impact of genomic profiling.
Genome Medicine,
2020, 12
Pubmed: 32907621
Direct link: https://doi.org/10.1186/s13073-020-00774-x
